



|
TOPICS BONOBO Chimpanzee "Ai" Crania photos Itani Jun'ichiro archives Open datasets for behavioral analysis Guidelines for Care and Use of Nonhuman Primates(pdf) Study material catalogue/database Guideline for field research of non-human primates 2019(pdf) Primate Genome DB 
Primate Research Institute, Kyoto University Copyright (c) |
Japanese Structural variations of subterminal satellite blocks and their source mechanisms as inferred from the meiotic configurations of chimpanzee chromosome termini
Hirohisa Hirai, Yuriko Hirai, Toshifumi Udono, Kiyoaki Matsubayashi, Anthony J. Tosi, Akihiko Koga
Abstract
African great apes have large subterminal satellite DNA (StSat) blocks in subtelomeric regions of the majority of their chromosomes, but humans lack these. Additionally, we have observed that the chimpanzee meiotic cell division process demonstrates unique partial terminal associations in pachytene. To examine the features of StSat blocks, we conducted molecular cytogenetic analyses undergoing mitotic metaphase (43 individuals) and meiotic cell division (one male). The results suggested that the subtelomeric associations are the mechanism for generating variability and genomic dispersion. Comparison between the present study and previous reports indicated that the chromosomal distribution rate of sutelomeric regions seems to have antagonistic correlation with arm numbers holding StSat blocks in humans, chimpanzees and gorillas. That is, the increase of StSat blocks probably reduces genomic diversity in the subtelomeric regions. The acquisition vs. loss of the StSat blocks is postulated as the underlying engine of this chromosomal differentiation yielded by meiotic chromosomal interaction. 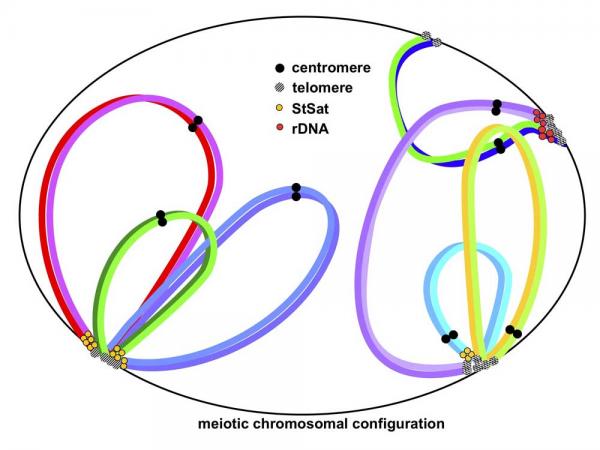 2019/08/19 Primate Research Institute
|