A novel composite retrotransposon derived from or generated independently of the SVA (SINE/VNTR/Alu) transposon has undergone proliferation in gibbon genomesHara T, Hirai Y, Baicharoen S, Hayakawa T, Hirai H, Koga A The superfamily Hominoidea (hominoids) comprises two families: Hominidae (hominids) and Hylobatidae (gibbons, also called small apes). The SVA transposon is a composite retrotransposon that occurs widely in hominoids and is considered to have been generated by stepwise fusions of three genetic elements: SINE-R, a variable number of tandem repeat (VNTR) sequence, and
Alu. We identified a novel transposon whose basic structure is the same as that of SVA, with one prominent difference being the presence of part of
prostaglandin reductase 2 (PTGR2) in place of SINE-R. We designate this composite transposon as PVA and propose two possible mechanisms regarding its generation. One is the derivation of PVA from SVA: the SINE-R region of SVA was replaced with
PTGR2 fragments by template switching. The other is the formation of PVA independently of SVA: a
PTGR2 fragment was fused to an evolutionary intermediate comprising the VNTR and Alu regions. The nucleotide sequence of the junction between the VNTR and
PTGR2 regions supports the second hypothesis. We identified PVA in the white-cheeked gibbon Nomascus leucogenys by analysis of genome sequence databases, and subsequent experimental analysis revealed its presence in all four gibbon genera. The white-cheeked gibbon harbors at least 93 PVA copies in its haploid genome. Another SVA-like composite transposon carrying parts of the LINE1 and
Alu transposons in place of SINE-R, designated as LAVA, has recently been reported. The significance of the discovery of PVA is that its substituted fragment originates not from a transposon but from a single-copy gene. PVA should provide additional insights into the transposition mechanism of this type of composite transposon; the transposition activity is conferred even if the substituted fragment is not related to a transposon. Genes & Genetic Systems 87 (3): 181-190. 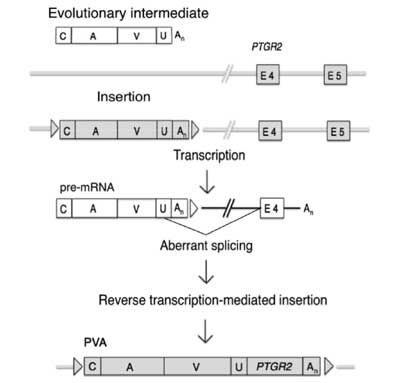
AUG/17/2012
Copyright(C) 2012 PRI ( ).
All rights reserved. ).
All rights reserved. |




